Tools
These methods enabled the discovery of diverse, potent compounds from uncultivated bacteria and poorly studied bacterial lineages.
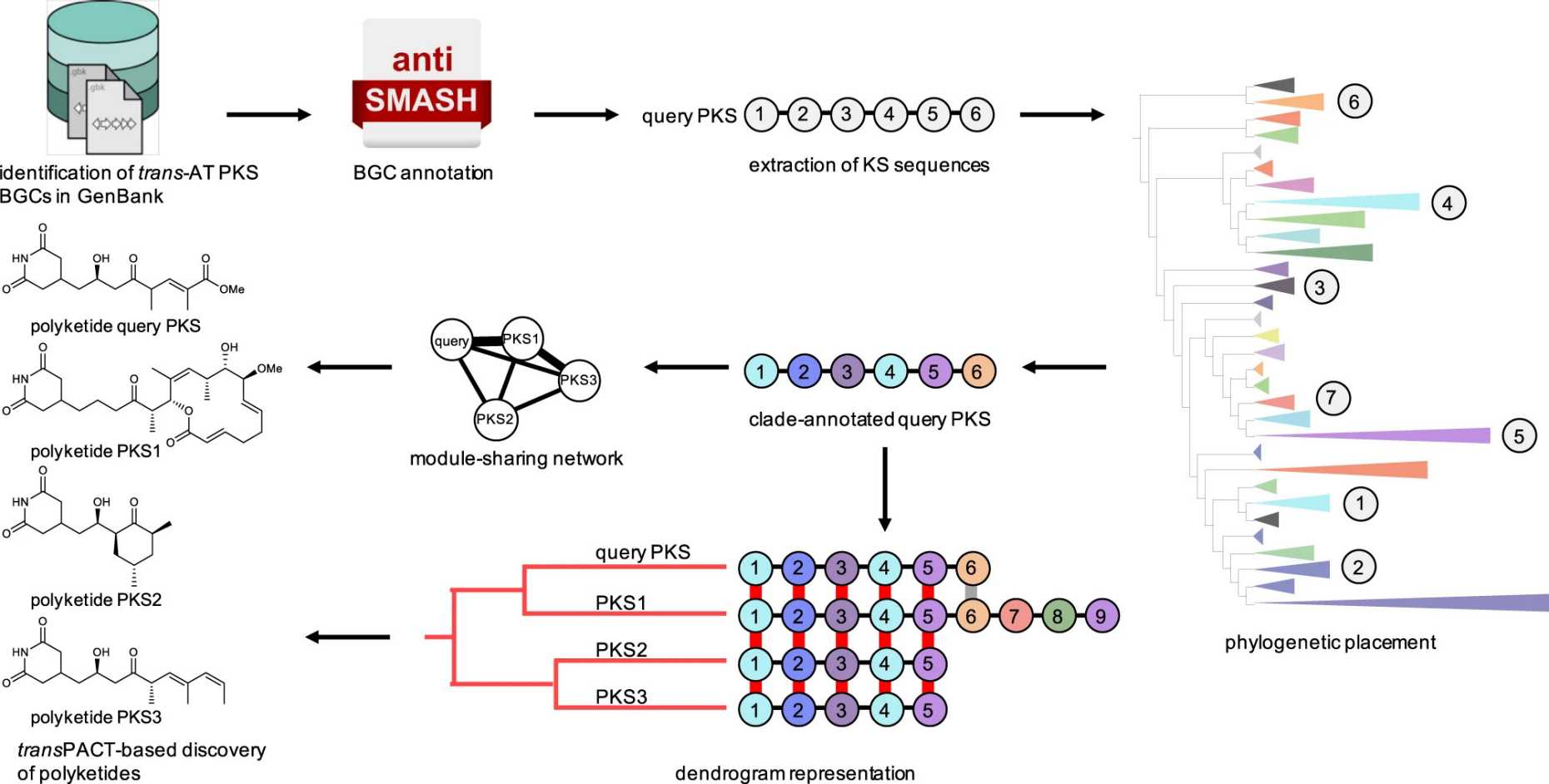
TransPACT
The external pagetransPACTcall_made algorithm (external pageNature Communicationscall_made) identifies conserved chemical substructures in trans-AT PKS-derived polyketides. We used it to study evolutionary patterns across bacterial genomes and to discover polyketide variants with conserved moieties.
- Code at external pageGitHubcall_made
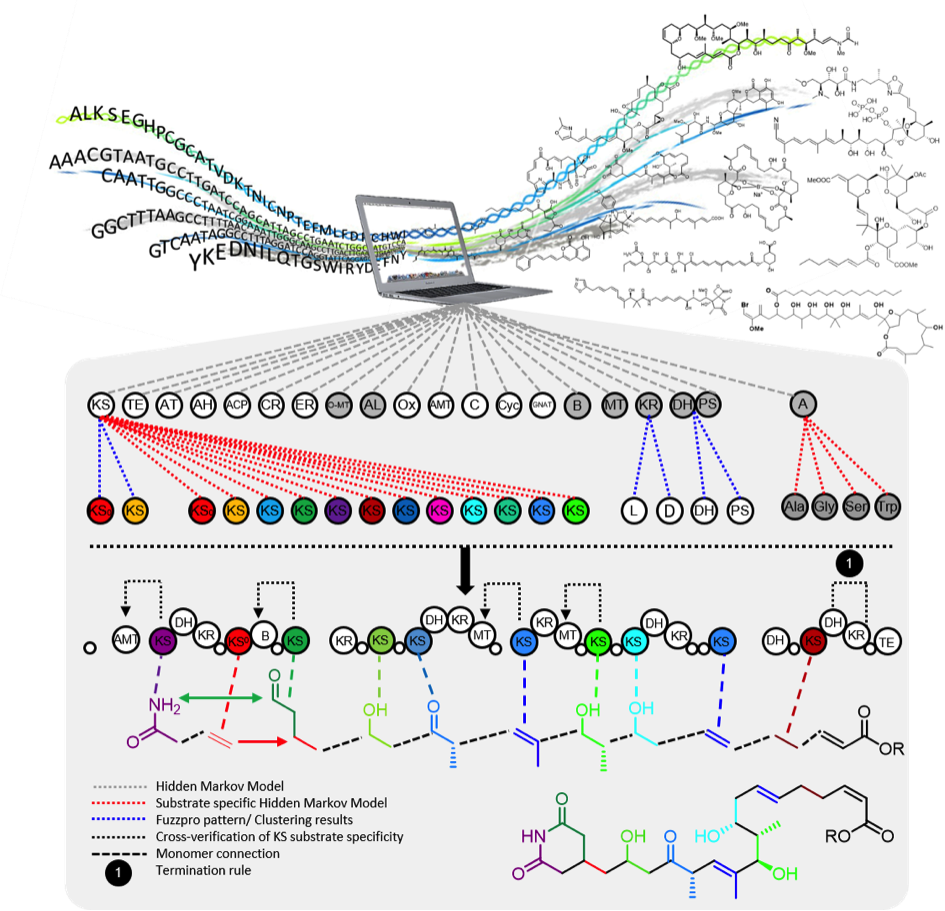
TransATor
The TransATor tool (external pageNature Chemical Biology)call_made makes structure predictions for trans-AT PKS-derived polyketides from protein sequences. We applied it for hypothesis-based polyketide discovery and biosynthetic studies.
- Free TransATor web application at transator.ethz.ch/trans-AT-PKS/
- Code and tutorial at external pageGitHubcall_made
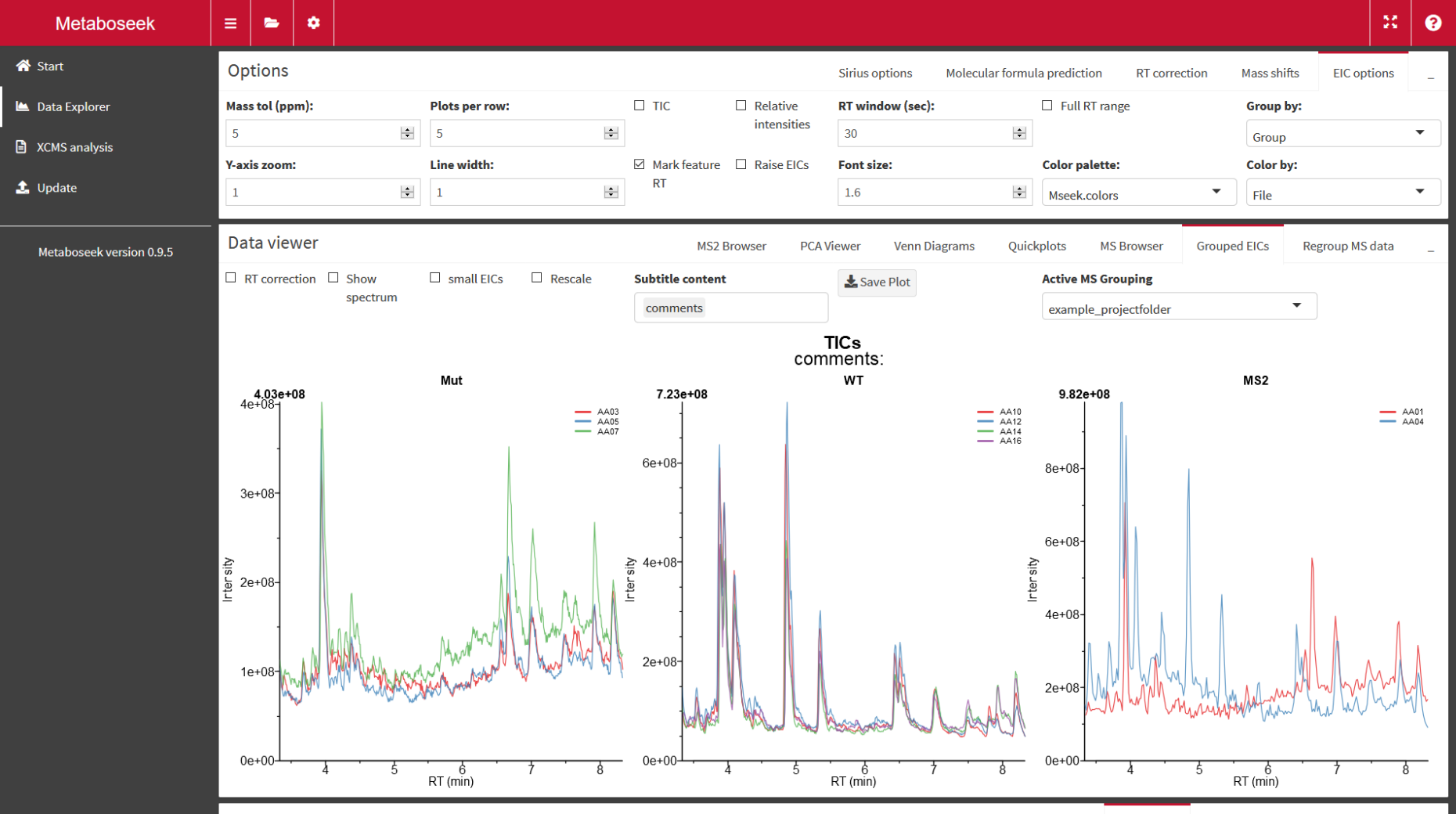
Metaboseek
external pageMetaboseekcall_made is a tool to analyze LC/MS data. Molecular features are detected and compared across mutliple samples. Metaboseek was developed by former PhD student Dr. Maximilian Helf at Cornell University (external pageNature Communicationscall_made).
- Free Metaboseek web application at external pagemetaboseek.comcall_made
- Code at external pageGitHubcall_made